tedana.decomposition.tedpca
- tedpca(data_cat, data_oc, mask, adaptive_mask, io_generator, tes, algorithm='aic', kdaw=10.0, rdaw=1.0, low_mem=False)[source]
Use principal components analysis (PCA) to identify and remove thermal noise from data.
- Parameters:
data_cat ((S x E x T) array_like) – Input functional data
data_oc ((S x T) array_like) – Optimally combined time series data
mask ((S,) array_like) – Boolean mask array
adaptive_mask ((S,) array_like) – Array where each value indicates the number of echoes with good signal for that voxel. This mask may be thresholded; for example, with values less than 3 set to 0. For more information on thresholding, see make_adaptive_mask.
io_generator (
tedana.io.OutputGenerator) – The output generation object for this workflowtes (
list) – List of echo times associated with data_cat, in millisecondsalgorithm ({‘kundu’, ‘kundu-stabilize’, ‘mdl’, ‘aic’, ‘kic’, float}, optional) – Method with which to select components in TEDPCA. PCA decomposition with the mdl, kic and aic options are based on a Moving Average (stationary Gaussian) process and are ordered from most to least aggressive (see [1]). If a float is provided, then it is assumed to represent percentage of variance explained (0-1) to retain from PCA. If an int is provided, then it is assumed to be the number of components to select Default is ‘aic’.
kdaw (
float, optional) – Dimensionality augmentation weight for Kappa calculations when algorithm is ‘kundu’. Must be a non-negative float, or -1 (a special value). Default is 10.rdaw (
float, optional) – Dimensionality augmentation weight for Rho calculations when algorithm is ‘kundu’. Must be a non-negative float, or -1 (a special value). Default is 1.low_mem (
bool, optional) – Whether to use incremental PCA (for low-memory systems) or not. This is only compatible with the “kundu” or “kundu-stabilize” algorithms. Default: False
- Returns:
kept_data ((S x T)
numpy.ndarray) – Dimensionally reduced optimally combined functional datan_components (
int) – Number of components retained from PCA decomposition
Notes
Notation
Meaning

Component pseudo-F statistic for TE-dependent (BOLD) model.
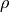
Component pseudo-F statistic for TE-independent (artifact) model.

Voxel
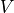
Total number of voxels in mask
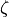
Something

Component
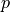
Something else
Steps:
Variance normalize either multi-echo or optimally combined data, depending on settings.
Decompose normalized data using PCA or SVD.
Compute
 and
and 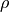 :
: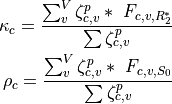
Some other stuff. Something about elbows.
Classify components as thermal noise if they meet both of the following criteria:
Nonsignificant
 and
and 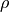 .
.Nonsignificant variance explained.
Generated Files
Default Filename
Content
desc-PCA_metrics.tsv
PCA component table
desc-PCA_metrics.json
Metadata sidecar file describing the component table
desc-PCA_mixing.tsv
PCA mixing matrix
desc-PCA_components.nii.gz
Component weight maps
desc-PCA_decomposition.json
Metadata sidecar file describing the PCA decomposition
References
See also
tedana.utils.make_adaptive_mask()The function used to create the
adaptive_maskparameter.tedana.constantsThe module describing the filenames for various naming conventions